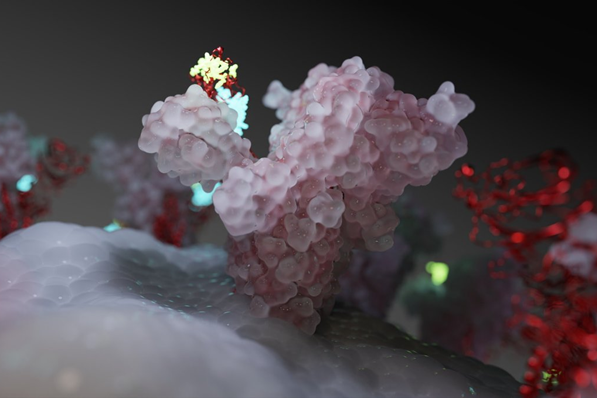
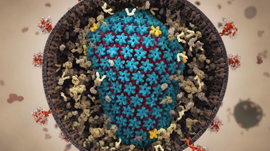
To help in the development of a treatment or vaccine for COVID-19, University of Texas at Austin and National Institute of Health (NIH) researchers recently achieved a critical breakthrough—creating the first 3D, atomic-scale map of the virus.
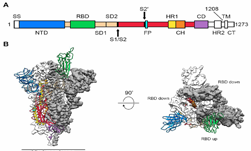
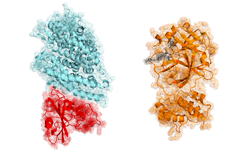
“2019-nCoV makes use of a densely glycosylated spike (S) protein to gain entry into host cells,” the researchers stated in a newly published Science paper, Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. “The CoV spike (S) glycoprotein is a key target for vaccines, therapeutic antibodies, and diagnostics.”
Jason McLellan, the associate professor at UT Austin who led the work, has spent years studying other coronaviruses, including SARS and MERS. The prior experience gave his team an advantage in identifying key features of the new virus, as noted in a recent post, Breakthrough in Coronavirus Research Results in New Map to Support Vaccine Design.
The new in-depth image relies on the Nobel Prize–winning cryo-electron microscopy, also called cryo-EM, to image the structure of the protein. To do this, the scientists prepared many copies of the virus molecule frozen in a thin layer of ice. The ice was then imaged in an electron microscope to create over 100,000 2D projection images of the molecule. The team then used GPU-accelerated software, cryoSPARC, to solve the 3D density image of the entire molecule.
CryoSPARC was developed by Toronto-based startup Structura Biotechnology, a company that grew out of a University of Toronto research project.
The application uses CUDA and CUDA libraries with NVIDIA V100 GPUs, and NVIDIA T4 GPUs on-premises and cloud service providers.
In total, it took McLellan and his team 12 days to go from a raw sample to producing the atomic-scale map.
The software can help academic researchers and pharmaceutical companies rapidly recover 3D protein structures, eliminating some of the guesswork out of the drug discovery process. The company says their software is in use by over 400 institutions, including the University of California, Berkeley, the Hospital for Sick Children, and other large structural biology labs.
“The structure of 2019-nCoV S should enable rapid development and evaluation of medical countermeasures to address the ongoing public health crisis,” the researchers explained.
For the next phase of the work, McLellan plans to use the molecule to isolate the naturally produced antibodies from patients who have been infected with the coronavirus and successfully recovered.
“In large enough quantities, these antibodies could help treat a coronavirus infection soon after exposure,” the researchers explained in the post.
To learn more about the technology behind this project, watch the on-demand webinar Cryo-EM for Drug Discovery, from Ali Punjani, CEO and co-founder of Structura Biotechnology.