To help tackle COVID-19, the long-running Folding@Home program, a distributed computing project for simulating protein dynamics, hit a breakthrough by achieving more than an exaflop of processing power. That’s more than 1,000,000,000,000,000,000 operations per second, all through crowdsourcing efforts.
For comparison, Summit, the world’s fastest supercomputer, which is powered by more than 27,000 NVIDIA V100 GPUs, can officially sustain around 150 petaflops, and 445 when benchmarked on HPL-AI.
In just a couple of days, nearly 400,000 gamers donated their GPU resources to build up the Folding@Home supercomputer.
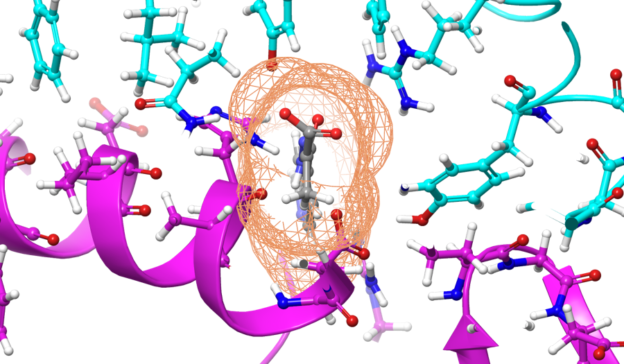
With this power, scientists aim to analyze the dynamics of the COVID-19 protein and hopefully gain better insights into potential drug interactions that can disable the virus.
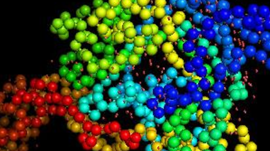
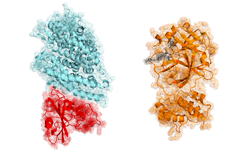
More specifically, a group of researchers at Washington University in St. Louis are tackling the problem of protein folding for COVID-19, the physical process by which a protein chain acquires its native three-dimensional structure.
“There are many experimental methods for determining protein structures. While extremely powerful, they only reveal a single snapshot of a protein’s usual shape. But proteins have lots of moving parts, so we really want to see the protein in action,” the organization stated in their web page, Coronavirus – What we’re doing and how you can help. “These calculations are enormous and every little bit helps! Each simulation you run is like buying a lottery ticket. The more tickets we buy, the better our chances of hitting the jackpot,” the organization said.
Folding@Home says they were the first distributed computing project to use GPUs for molecular dynamics simulations.
“For certain types of calculations, we’ve seen GPUs give us a 20-30x speedup over their CPU-based counterparts,” the organization stated.
To donate your GPU resources, download the Folding@home software.
(Originally published on March 30, 2020)