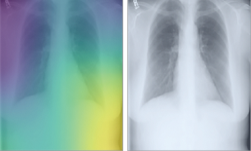
A team of Stanford researchers developed a deep learning-based algorithm that evaluates chest X-rays for signs of disease.
“Interpreting X-ray images to diagnose pathologies like pneumonia is very challenging, and we know that there’s a lot of variability in the diagnoses radiologists arrive at,” said Pranav Rajpurkar, a graduate student in the Stanford Machine Learning Group and co-lead author of the paper. “We became interested in developing machine learning algorithms that could learn from hundreds of thousands of chest X-ray diagnoses and make accurate diagnoses.”
Using GTX 1080s and TITAN X GPUs with the cuDNN-accelerated PyTorch deep learning framework, the researchers trained their model CheXNet (a 121-layer convolutional neural network) on the ChestX-ray14 dataset that consists of over 100,000 frontal-view X-ray images with 14 different thoracic diseases, including pneumonia.
According to the researchers, detecting pneumonia in chest radiography can be difficult for radiologists since the appearance of pneumonia in X-ray images is often vague, can overlap with other diagnoses, and can mimic many other benign abnormalities. These discrepancies cause considerable variability among radiologists in the diagnosis of pneumonia.
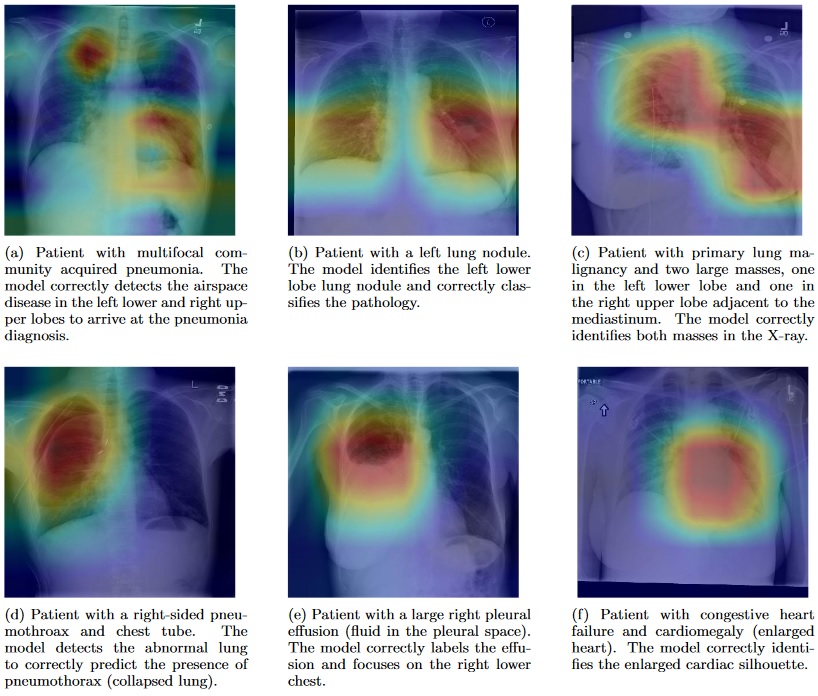
Within a week the researchers had an algorithm that diagnosed 10 of the pathologies labeled in the X-rays more accurately than previous state-of-the-art results. In just over a month, their algorithm could beat these standards in all 14 identification tasks. In that short time span, CheXNet also outperformed the four Stanford radiologists in diagnosing pneumonia accurately.
“We plan to continue building and improving upon medical algorithms that can automatically detect abnormalities and we hope to make high-quality, anonymized medical datasets publicly available for others to work on similar problems,” said Jeremy Irvin, a graduate student in the Machine Learning group and co-lead author of the paper. “There is massive potential for machine learning to improve the current health care system, and we want to continue to be at the forefront of innovation in the field.”
Read more >
Algorithm Successfully Diagnoses Pneumonia at Radiologist-Level Accuracy
Dec 12, 2017
Discuss (0)
Related resources
- GTC session: Early Diagnosis of Cancer Cachexia Using Body Composition Index as the Radiographic Biomarker
- GTC session: Creating AI-Powered Hardware Solutions for Medical Imaging Applications
- GTC session: Optimizing Your AI Strategy to Develop and Deploy Novel Deep Learning Models in the Cloud for Medical Image Analysis
- NGC Containers: Microvolution
- SDK: MONAI Cloud API
- SDK: Clara Viz