Artificial intelligence is revolutionizing how medical images are interpreted, helping medical professionals save time analyzing magnetic resonance imaging, CT scans, and X-rays. However, one of the significant challenges deep learning scientists working in the medical community face is the lack of accurate and reliable data to train their neural networks.
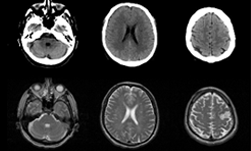
For the first time, researchers are now using GANs to generate abnormal brain MRIs that can be used to train neural networks. To solve this problem a team of researchers from NVIDIA, the Mayo Clinic, and the MGH & BWH Center for Clinical Data Science developed a deep learning-based model that can generate accurate and reliable synthetic images that can be used for training an artificial intelligence system.
“Data diversity is critical to success when training deep learning models,” the researchers stated in their paper. “Medical imaging data sets are often imbalanced as pathologic findings are generally rare, which introduces significant challenges when training deep learning models. We propose a method to generate synthetic abnormal MRI images with brain tumors by training a generative adversarial network.”
Using an NVIDIA DGX-system, which contains NVIDIA Tesla V100 GPUs, with the cuDNN-accelerated PyTorch deep learning framework the team trained their generative adversarial network on data from two publicly available datasets of brain MRIs. One dataset contains thousands of 3D T1-weighted brain MRIs with Alzheimer’s disease, the other contains about two hundred 4D brain MRIs with brain tumors.
The team based their image-to-image translation method on the pix2pix model, previously developed by NVIDIA researchers.
GANs have been used in medical imaging before to generate a motion model from a single preoperative MRI, upsample a low-resolution image, create a synthetic head CT from a brain MRI, perform medical segmentation, and automatically align different types of MRIs, saving doctors hours.
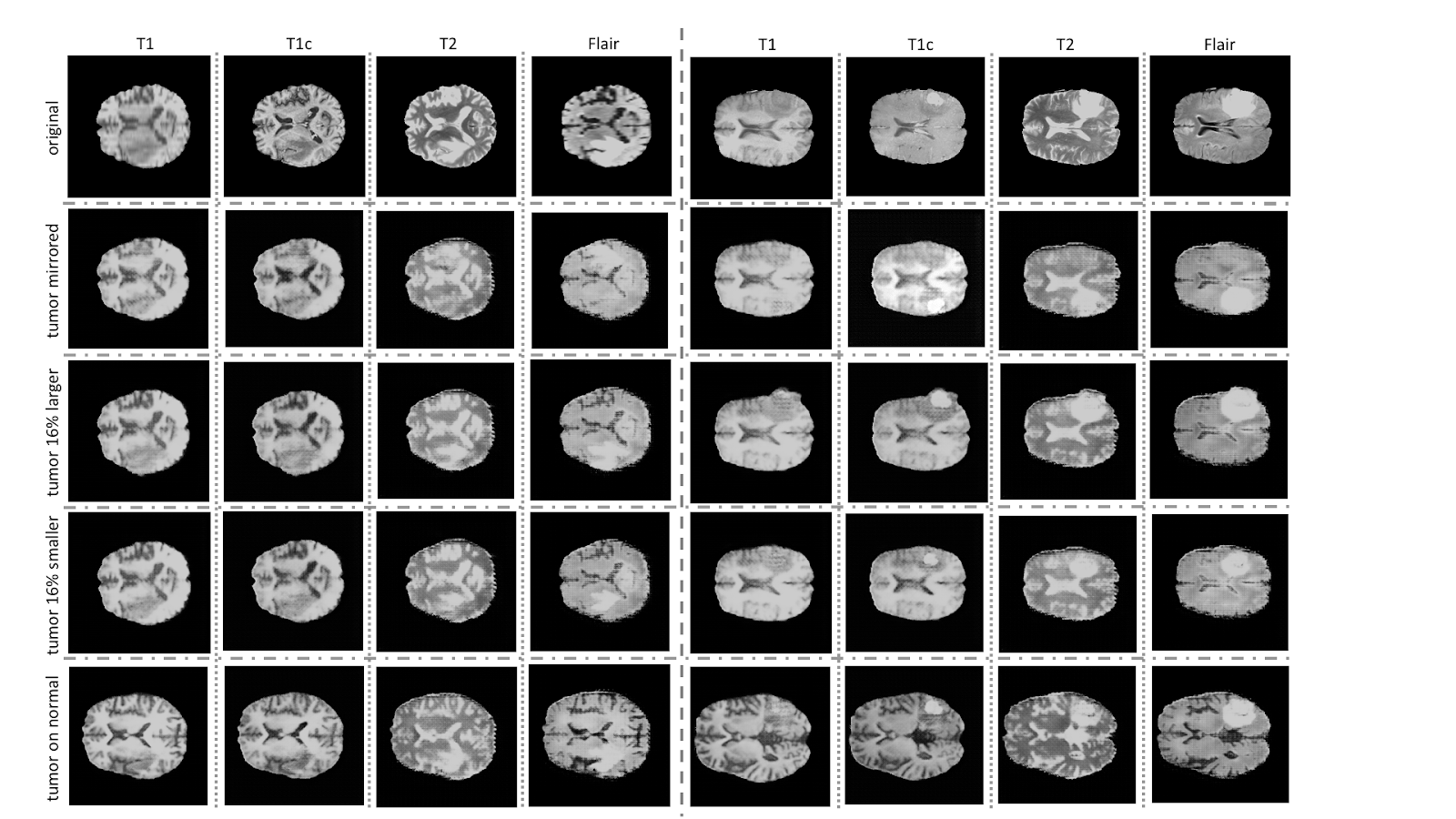
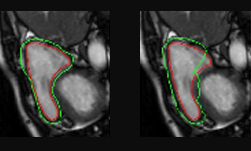
“This offers an automatable, low-cost source of diverse data that can be used to supplement the training set,” the researchers stated. “ For example, we can alter a tumor’s size, change its location, or place a tumor in an otherwise healthy brain, to systematically have the image and the corresponding annotation,” the team explained.
Because the images are synthetically generated, there is no patient data or privacy concerns. Medical institutions can easily share data they generate with other institutions, creating millions of different combinations that can be used to accelerate the work.
The team hopes that the model can immediately help deep learning scientists generate new data that can be used to detect abnormalities and in the end, save lives.
The researchers published their paper on ArXiv.
Read more>
AI Can Generate Synthetic MRIs to Advance Medical Research
Sep 16, 2018
Discuss (0)
Related resources
- DLI course: Data Augmentation and Segmentation with Generative Networks for Medical Imaging
- GTC session: AI-Enabled MR Reconstruction: Doing More With Less
- GTC session: Deep Learning-Based GRAPPA Operator Gridding for Non-Cartesian MR Image Reconstruction
- GTC session: Revolutionizing Healthcare through AI-Empowered Solutions and Medical Devices
- SDK: MONAI Cloud API
- Webinar: Isaac Developer Meetup #2 - Build AI-Powered Robots with NVIDIA Isaac Replicator and NVIDIA TAO