‘Meet the Researcher’ is a monthly series in which we spotlight different researchers in academia who are using NVIDIA technology to accelerate their work. This month, we spotlight Gregory A. Voth, Distinguished Professor at the Department of Chemistry, The University of Chicago.
Voth received a Ph.D. in Theoretical Chemistry from the California Institute of Technology in 1987 and was an IBM Postdoctoral Fellow at the University of California, Berkeley from 1987-89. Voth has authored or co-authored more than 550 peer-reviewed scientific articles that have been cited approximately 44,000 times.
What are your research areas of focus?
I have led the development and application of theoretical and computational methods to study problems involving the structure and dynamics of complex condensed phase systems, including proteins, membranes, liquids, and materials. And pioneered a method known as “multiscale coarse-graining” in which the resolution of the molecular-scale entities is reduced into simpler structures, but key information on their interactions is accurately retained (or renormalized) so the resulting computer simulation can accurately and efficiently predict the properties of large assemblies of complex molecules such as lipids and proteins. This method is multiscale, meaning it describes complex condensed phase and biomolecular systems from the molecular scale to the mesoscale and ultimately to the macroscopic scale.
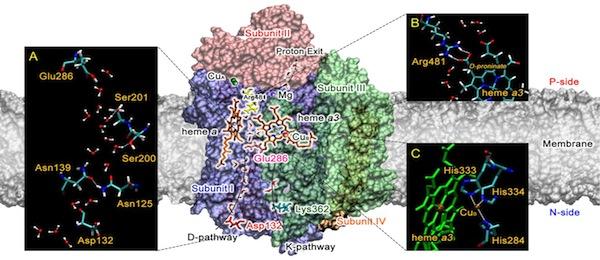
My other research interests include the study of charge transport (protons and electrons) in water and biomolecules – a fundamental process in living organisms and other systems that has been poorly understood because of its complexity. I also study the exotic behavior of room-temperature ionic liquids and other complex materials such as nanoparticle self-assembly, polymer electrolyte membranes for fuel cells, and electrode-electrolyte interfaces in energy storage devices. In the earlier part of my career, I extensively developed and applied new methods to study quantum and electron transfer dynamics in condensed phase systems-much of this work was based on the Feynman path integral description of quantum mechanics.
What motivated you to pursue this work?
I have always been focused on understanding the complex problems underpinning realistic scientific experiments, such as those uncovering the mechanisms of proteins and cells. Developing multiscale theories and methods that allow computational scientists such as myself to study and understand these problems has only strengthened my interest in attacking real world problems.
What problems or challenges does your research address?
My research addresses challenges in the theory and simulation of complex chemical systems at multiple scales. An underlying theme of my work is the development of new computational methods that enable simulations that cannot be performed using standard techniques. In particular, I gravitate towards problems where fundamental knowledge of collective dynamical behavior is lacking.
What is the (expected) impact of your work on the field/community/world?
My work has broadly impacted the scientific community. In particular, my group has pioneered the theory and design of multiscale algorithms to investigate complex condensed phase and biomolecular systems. Our methodological capabilities encompass enhanced free energy techniques to aid in rare event sampling, systematic coarse-graining to derive molecular models with reduced resolution, and the development of open-source software that implements these algorithms and disseminates their use to the wider scientific community.
How have you used NVIDIA technology either in your current or previous research?
I am using NVIDIA GPUs to accelerate various aspects of my computational research. For instance, my group is developing a GPU-enabled plug-in for open-source MD engines that will allow us and other researchers to study charge transport using our multiscale reactive molecular dynamics (MS-RMD) method; we expect GPUs to provide at least an order of magnitude speed up in performance. My research interests also include biomolecules that undergo slow and collective conformational transitions. The use of CUDA-accelerated MD engines, in combination with my group’s enhanced sampling algorithms and coarse-graining methodologies, have allowed us to efficiently investigate these systems without the need to scale to hundreds of CPU nodes. Most recently, this was performed to probe the molecular dynamics of SARS-CoV-2 viral proteins at the DOE’s ORNL Summit.
Tell us about your recent breakthroughs?
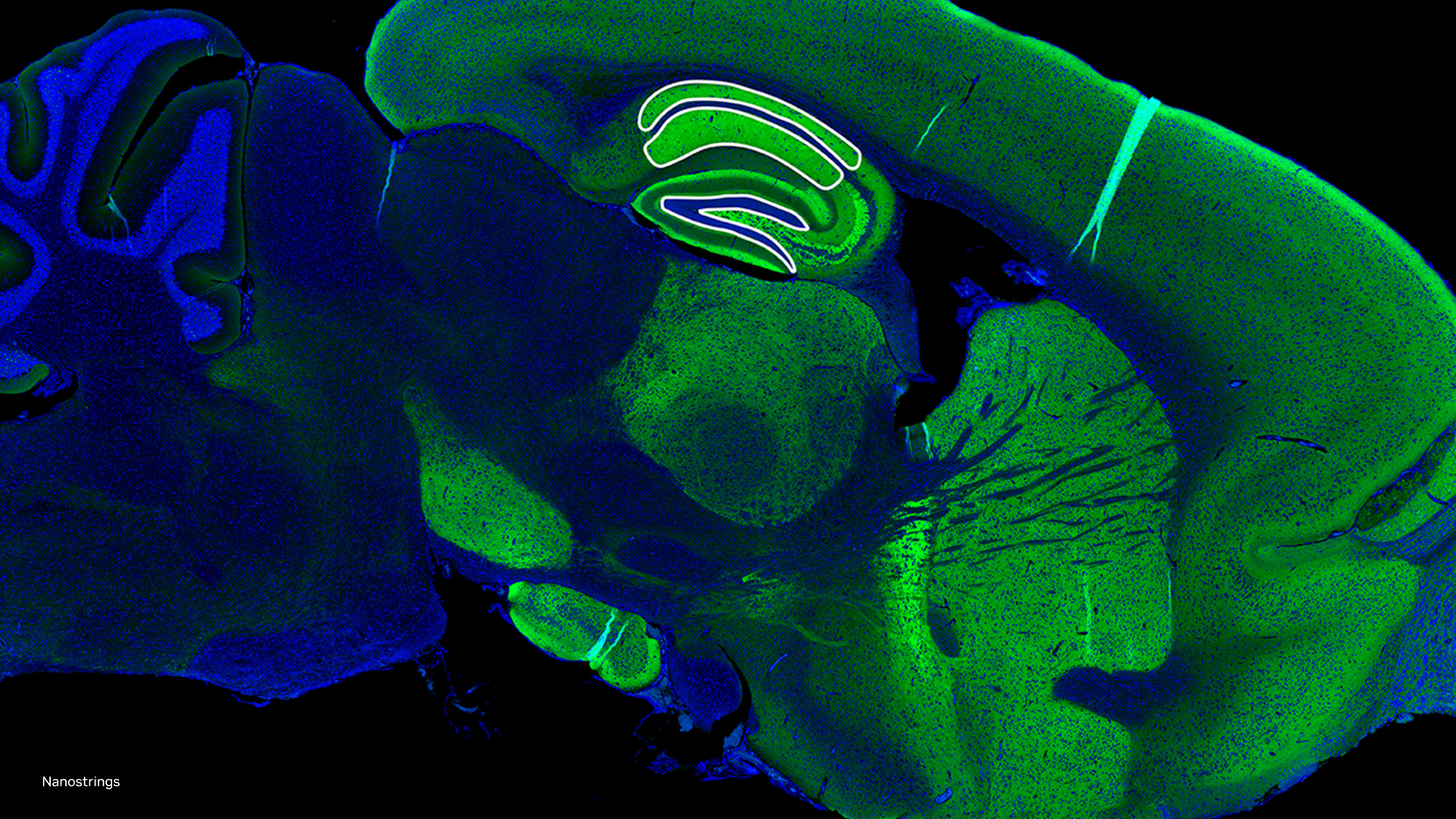
Recently, NVIDIA GPUs enabled us to characterize the dynamics of essential coronavirus proteins. Our preliminary findings reveal collective dynamics of the spikes that decorate coronavirus particles; these spikes mediate viral particle targeting and binding to new cells during infection. Expect more publications soon that will showcase our use of NVIDIA technology.

What’s next for your research?
We will continue to work at the interface of theory and experiment, using computational methods to bridge the gap in simulation scales for macromolecular behavior. An exciting frontier is to develop and apply ideas from the machine-learning community to push the boundaries of multiscale modeling and simulation. The current COVID-19 pandemic has also made it clear that we need better tools to rapidly integrate data from a multitude of sources (including MD and experimental data), autonomously fit multiscale models, and iteratively improving model parameterization. An important goal of my future research is to develop these methods and establish an integrated computational pipeline to help scientists accelerate their own research.
Any advice for new researchers?
Never give up! Be brave! Believe that what you are doing is important. And do not be afraid to ask for input or advice from your experimentalist colleagues.