Using supercomputers equipped with NVIDIA Tesla GPUs, a team of researchers at Colorado State University have identified a critical protein structure in the dengue virus that could potentially prevent the replication of the disease.
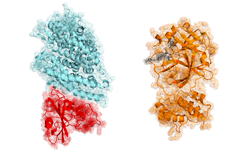
The study, recently published in the PLOS Computational Biology Journal, focused on a small segment of a viral enzyme called nonstructural protein 3 (NS3). The enzyme plays a crucial role in the conditions dengue requires to survive and spread.
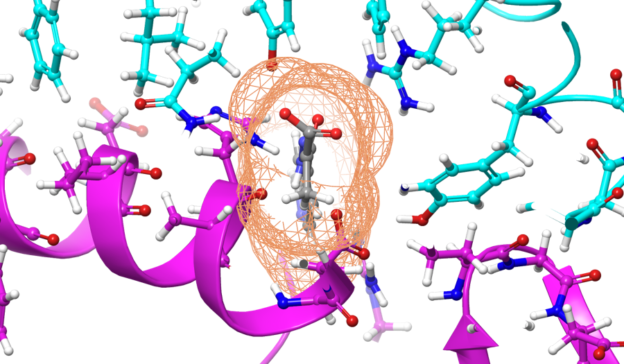
“Our analyses identify a previously underappreciated part of the NS3 enzyme known as ‘motif V’ that serves as a communication hub between two critical binding sites needed for RNA replication,” Martin McCullagh, assistant professor of chemistry at Colorado State University and the study’s principal researcher said in a press release. “Our results suggest that this hub could be a novel target for NS3 inhibitors.”
For decades, researchers all over the world have looked into the possibility of using NS3 to develop drugs capable of preventing the replication of the dengue virus. However, many scientists worry the NS3 protein sequence is too similar to human proteins, which could create unwanted side effects.
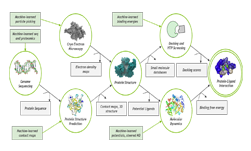
To better understand the enzyme, the Colorado State University researchers developed a model that sheds more light on how the NS3 works on a molecular scale. Using San Diego Supercomputing Center’s Comet supercomputer and Bridges supercomputer at the Pittsburgh Supercomputing Center, both equipped with NVIDIA Tesla GPUs, the team stretched the amount of time proteins are simulated in their natural state by 12.5 microseconds, 100 times longer than previously reported simulations on dengue NS3.
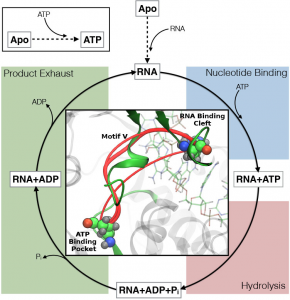
The extra time, though small, revealed the previously unknown role played by motif V in the NS3 enzyme.
The researchers say they will use the model to study the Zika and West Nile viruses and how the NS3 enzyme behaves in those diseases. The study has the potential to help scientists develop drugs that can one day stop the diseases from spreading.
Read more >
GPU-Accelerated Supercomputers Aid Researchers Working to Stop the Dengue Virus
Apr 30, 2018
Discuss (0)

Related resources
- DLI course: Speed Up DataFrame Operations With RAPIDS cuDF
- GTC session: Poster Reception (Sponsored by Cadence)
- GTC session: Priming Researchers and Students for AI and Accelerated Computing Breakthroughs With Self-Sustaining Training Programs
- GTC session: RAPIDS Accelerator for Apache Spark Propels Data Center Efficiency and Cost Savings
- NGC Containers: CUDA
- SDK: cuSOLVER