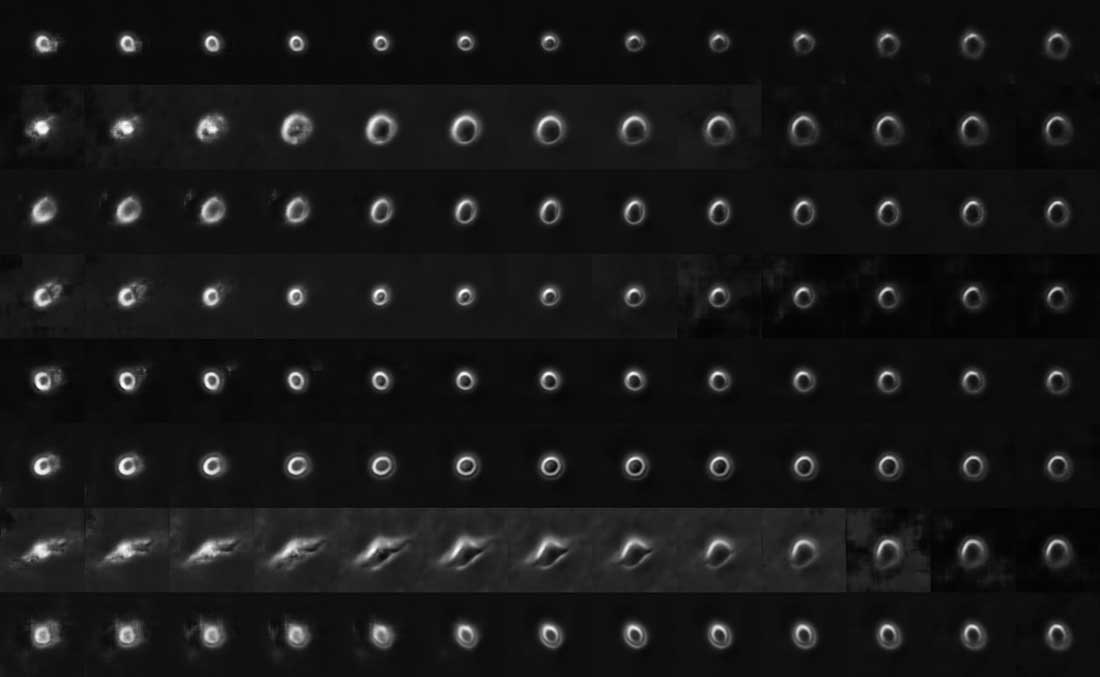
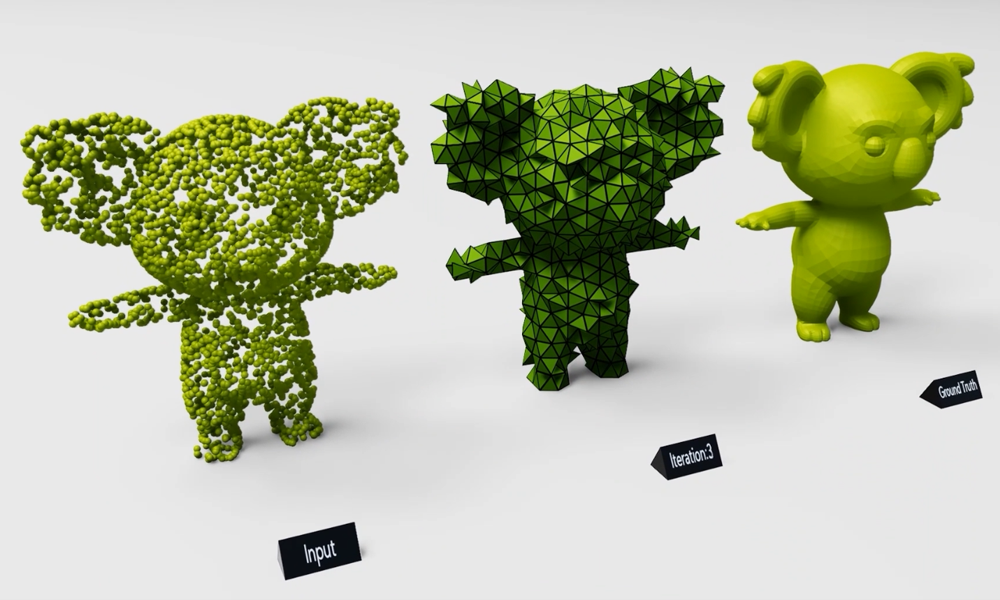
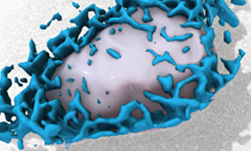
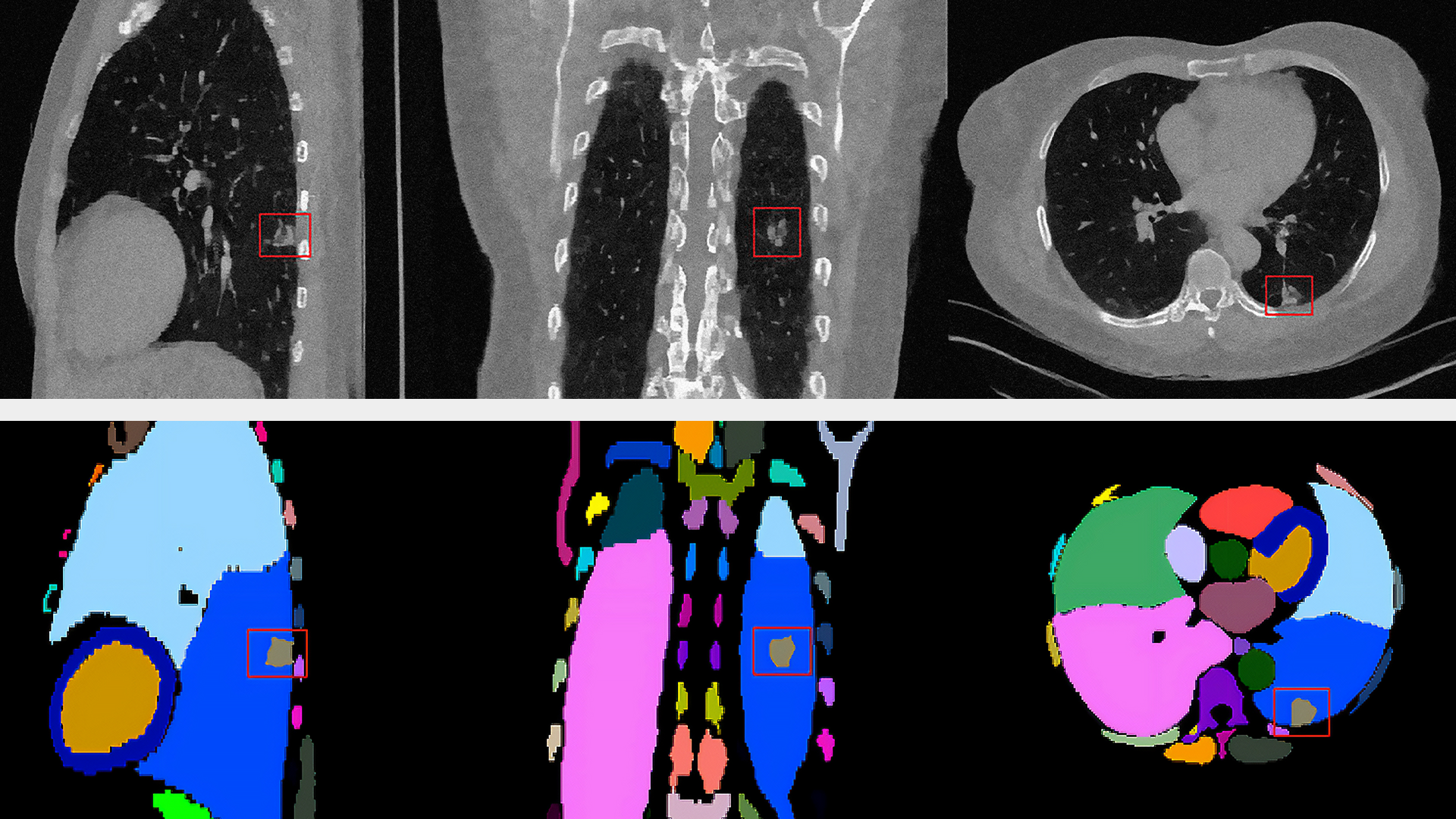
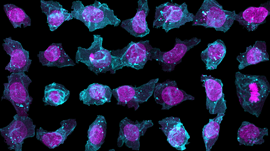
The Allen Institute for Cell Science launched a one-of-a-kind online portal of 3D cell images called Allen Cell Explorer that were produced using deep learning.
The website combines large-scale 3D imaging data, the first application of deep learning to create predictive models of cell organization, and a growing suite of powerful tools.
“This is the first time researchers have used deep learning to try and understand the elusive question of how actual cells are organized,” says Rick Horwitz, Ph.D., Executive Director of the Allen Institute for Cell Science. “The cartoons we rely on in textbooks, which are based on an artist’s interpretation of data from a relatively small number of cells, will eventually be replaced by data driven models of this kind from very large numbers of cells.”
Using TITAN X Pascal GPUs on NVIDIA Docker with cuDNN, the scientists trained their deep learning models on over six thousand 3D pluripotent human cells and found relationships between the locations of cellular structures. They then used that information to predict where the structures might be when the program was given just a couple of clues, such as the position of the nucleus. The program ‘learned’ by comparing its predictions to actual cells.
“Cells are incredibly complex, with thousands of moving and interacting parts that work together to drive and regulate both cell architecture and behavior,” says Horwitz. “We are beyond excited to launch the Allen Cell Explorer website and to share our cells, incredible image data, predictive models and more with the global scientific community.”
The 3D interactive tool should go live later this year. Currently, the site shows a preview of how it will work using side-by-side comparisons of predicted and actual images.
Read more >
Deep Learning Predicts the Look of Cells
Apr 07, 2017
Discuss (0)
Related resources
- DLI course: Deep Learning for Industrial Inspection
- GTC session: Computer Vision for Rare Disease Genomic Medicine
- GTC session: How Artificial Intelligence is Powering the Future of Biomedicine
- GTC session: How AI Will Decode Biology to Radically Improve Lives
- NGC Containers: MATLAB
- SDK: Neural VDB