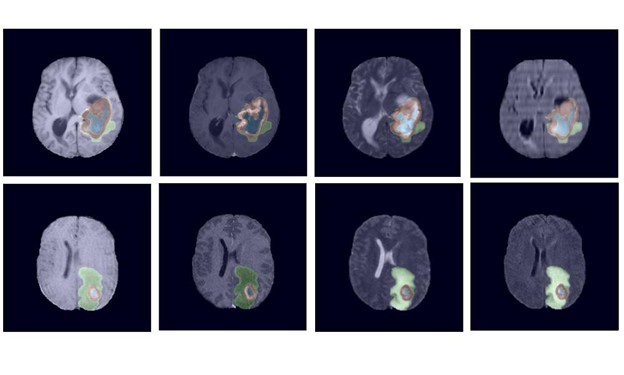
Each year tens of thousands of people in the United States are diagnosed with a brain tumor. To help physicians more effectively analyze, treat, and monitor tumors, NVIDIA researchers have developed a robust deep learning-based technique that uses 3D magnetic resonance images to automatically segment tumors. Segmentation provides tumor boundary definition of the affected region.
In countries with a shortage of trained experts, the technology could one day serve as a life-saving tool that helps patients receive the care they need.
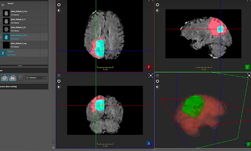
ImFusion visualization software helped the team run inference and visualize the results directly from the GUI, with easy configuration setup, the team said.
“Automated segmentation of 3D brain tumors can save physicians time and provide an accurate reproducible solution for further tumor analysis and monitoring. In this work, we describe our semantic segmentation approach for volumetric 3D brain tumor segmentation from multimodal 3D MRIs, which won the BraTS 2018 challenge,” said Andriy Myronenko, a senior research scientist at NVIDIA.
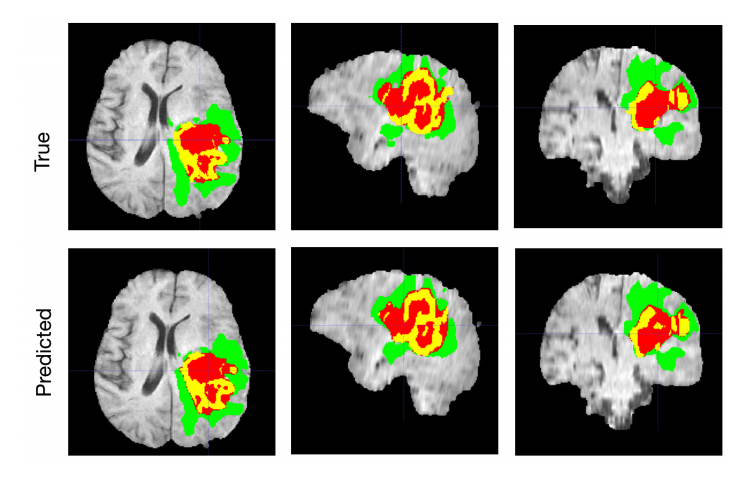
The BraTS challenge, or the Multimodal Brain Tumor Segmentation Challenge, is an international competition focused on the segmentation of brain tumors. The challenge is organized by the University of Pennsylvania’s Perelman School of Medicine.
In developing the work, Myronenko focused on gliomas, one of the most common types of primary brain tumors. High-grade gliomas are an aggressive type of malignant brain tumor and the best tools physicians have to diagnose them are magnetic resonance images. However, the process of manually delineating an image, or determining the exact position of a border or boundary of the tumor requires anatomical expertise. The process is also expensive, and prone to human error. That is why automatic segmentation is such an important tool.
Using data from 19 institutions and several MRI scanners, Myronenko trained an encoder-decoder convolutional neural network to extract the features of a brain MRI.
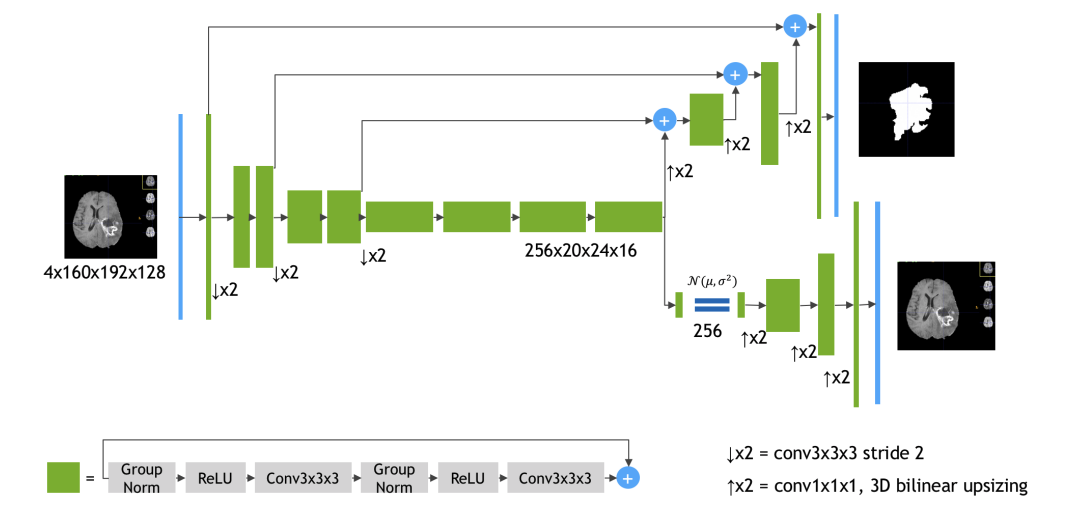
The encoder extracts the features of the images and the decoder reconstructs the dense segmentation masks of an image.
The network was trained on NVIDIA Tesla V100 GPUs and on a DGX-1 server with the cuDNN-accelerated TensorFlow deep learning framework. “During training we used a random crop of size 160x192x128, which ensures that most image content remains within the crop area,” Myronenko said. “
The BraTS 2018 testing dataset results are 0.7664, 0.8839 and 0.8154 average dice for enhanced tumor core, whole tumor and tumor core, respectively, Myronenko wrote in the paper.
The research is being presented at RSNA’s 104th Scientific and Annual Meeting in Chicago, Illinois.
Read more>
Automatically Segmenting Brain Tumors with AI
Nov 27, 2018
Discuss (1)
Related resources
- DLI course: Data Augmentation and Segmentation with Generative Networks for Medical Imaging
- DLI course: Image Classification with TensorFlow: Radiomics?€?1p19q Chromosome Status Classification
- DLI course: Coarse-to-Fine Contextual Memory for Medical Imaging
- GTC session: Semantic Segmentation of Meningioma Tumor Subregions on Multiparametric 3D MRI using Adaptive Convolutional Neural Networks
- GTC session: Edge-Enhanced Ensemble Learning Approach for Super-Resolution of T2-Weighted Brain MR Images
- GTC session: Optimizing Your AI Strategy to Develop and Deploy Novel Deep Learning Models in the Cloud for Medical Image Analysis